Gene Ontology
The discipline of Gene Ontology is a bioinformatic approach to present various attributes of a particular gene by unifying the genes 3 main biological domains: the cellular component, biological function, and biological process. These gene product properties allow for functional interpretation of experimental data [1]. Various tools are available for the gathering data from gene domains coming from various programs such as the GO project, AmiGO, & OBO-edit [3].
The discipline of Gene Ontology is a bioinformatic approach to present various attributes of a particular gene by unifying the genes 3 main biological domains: the cellular component, biological function, and biological process. These gene product properties allow for functional interpretation of experimental data [1]. Various tools are available for the gathering data from gene domains coming from various programs such as the GO project, AmiGO, & OBO-edit [3].
|
Cellular Component
This domain describes a location, relative to to other cellular components and structures. The location of gene products are described in one of two ways: relative to other cellular components or the stable macromolecular complexes they’re a part of. The domain distinguishes itself from the other two domains as this refers to the cellular anatomy [2].
|
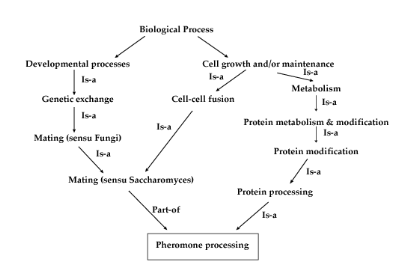
Biological Process
This domain describes a series of events accomplished by one or more assemblies of molecular functions. The distinguishing factor between molecular function and biological process is that this domain requires more than one step [1].
|
Molecular Function
This domain describes the activities performed by the gene product at the molecular level E.g) Transcription regulator activity, zinc ion binding, or glucose transporter activity [8].
|
ADA Isoforms
|
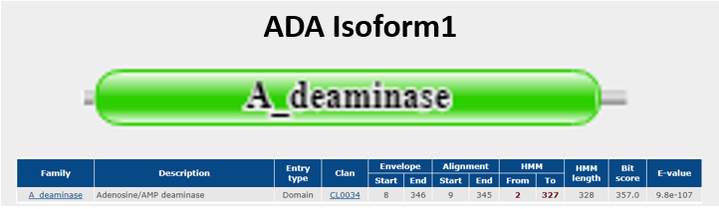
Figure 1a shows the protein domain found in ADA Isoform 1 one that was searched and gethered from Pfam [13]. Isoform 1 does not show any specific domain interactions.
|
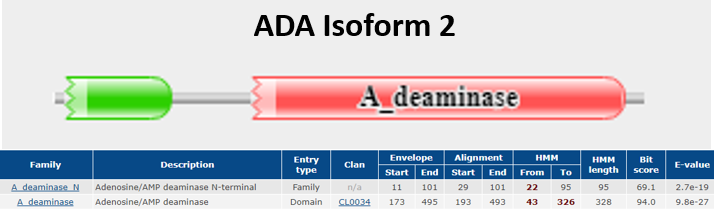
Figure 1b shows the protein domain found in ADA Isoform 2 collected from Pfam [13]. Aside from the Adenosine/AMP deaminase also found in Isoform 1, it also demonstrates interaction with Adenosine/AMP deaminase N-terminal. [13].
|
ADA Ontologies
|
Cellular Component
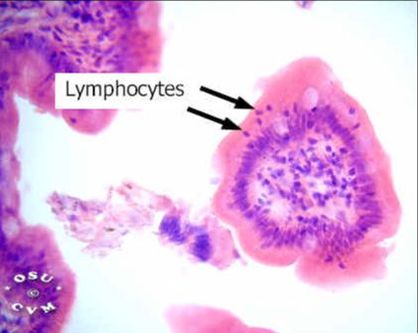
ADA is found all over the human body but is predominantly expressed in cytoplasm of lymphocyte cells and macrophages[5]. Since the enzyme is ubiquitously expressed, nonhematopoietic tissues such as kidneys, bones, cartilage and the central nervous system are also affected by ADA deficiency [4]. There exists two human isoforms of Adenosine Deaminase: ADA1 and ADA2 [6]. ADA1 & 2’s location is conserved in macrophages but they differentiate in many other places. |
Figure [A](left) showing a micrograph image of free lymphocytes found in lamina epithelialis and lamina propria in a small intestine.
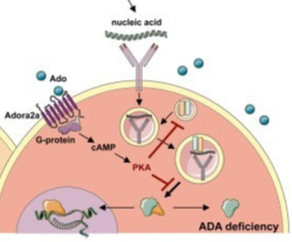
Figure [B] (right) showing the cellular component of ADA found in the cytoplasm of a lymphocyte cell. |
|
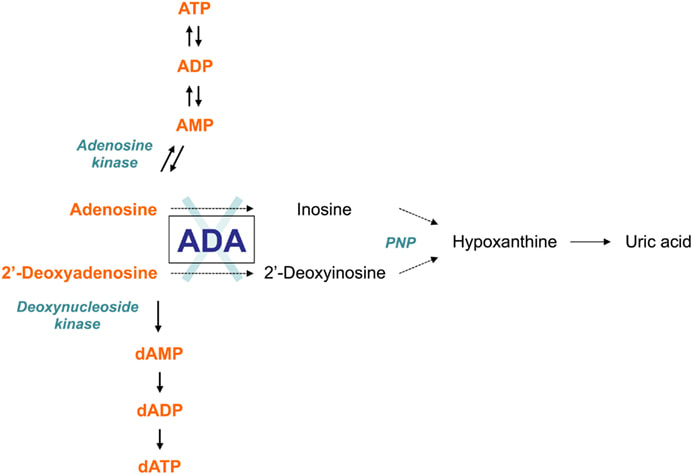
This domain describes a series of events accomplished by one or more assemblies of molecular functions. The distinguishing factor between molecular function and biological process is that this domain requires more than one step [1].ADA is mainly involved in Purine metabolism as well as development and maintenance of the immune system. ADA also has associations with epithelial cell differentiation, neurotransmission, and gestation maintenance [7].
|
|
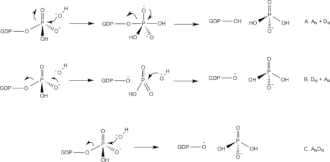
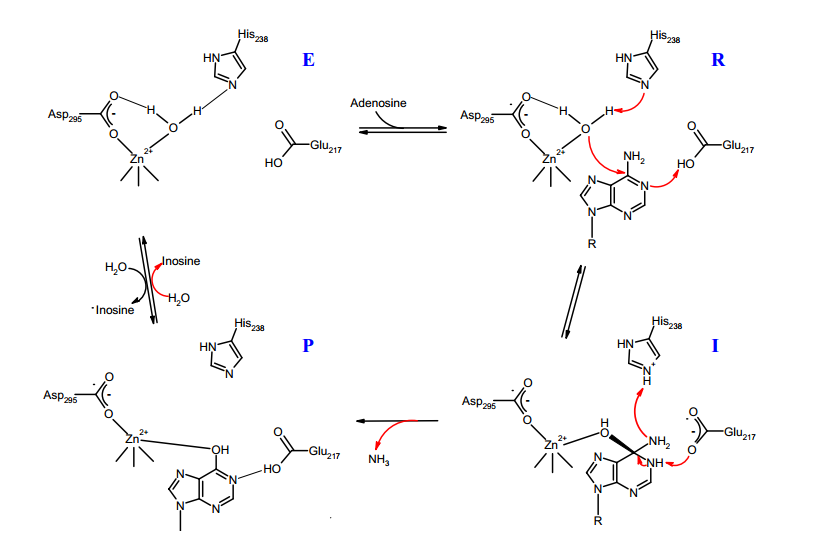
This domain describes the activities performed by the gene product at the molecular level E.g) Transcription regulator activity, zinc ion binding, or glucose transporter activity [8]. Adenosine Deaminase is involved in an irreversible deamination of of adenosine through an Sn2 reaction that substitutes the amino group with a ketone [9]. By either mechanism, Zn2+ as a strong electrophile activates a water molecule, which is deprotonated by the basic Asp295 to form the attacking hydroxide[10]. His238 orients the water molecule and stabilizes the charge of the attacking hydroxide. Glu217 is protonated to donate a proton to N1 of the substrate. The reaction is stereospecific due to the location of the Zinc, Asp295, and His238 residues, which all face the B-side of the purine ring of the substrate [10].
|
Figure [C] shows a simplified version of the deamination reaction ocurring between adenosine deaminase and adenosine thorugh the functional side chain amino acids of the ADA enzyme working together to remove the amino group of Adenosine and replace it with a ketone group.
|
|
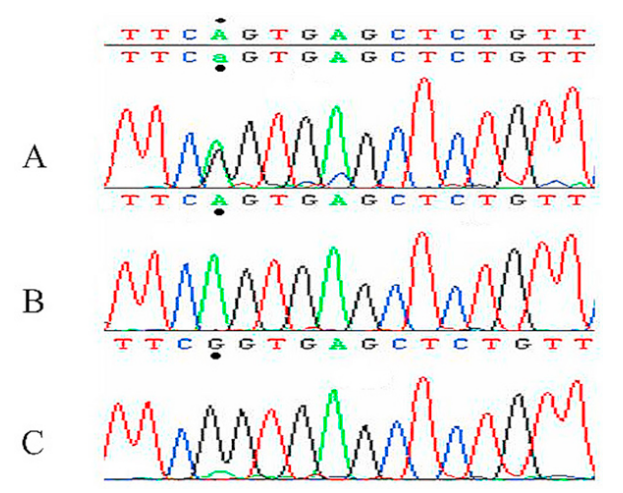
Figure D shows a chromatograph presenting the segment of Exon 9 with the observed G>A mutation. Child patient B is an affected carrier of the mutation as shown with the marked amino acid (a.a) being Glutamine as opposed to his unaffected sister showing the Arginine a.a [12].
|
Location of ADA Mutation
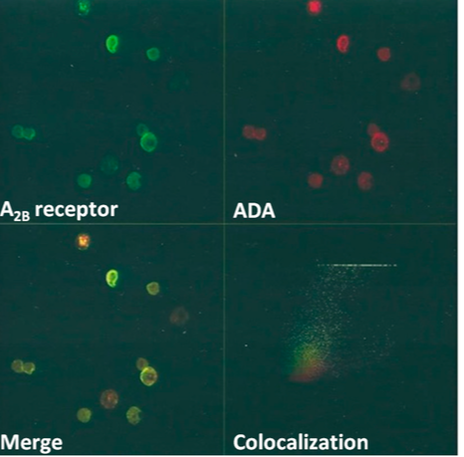
Previous studies on SCID patients have indicated a few different point mutations responsible for ADA deficiency with m amino acid substitutions and other alternative splicing results have demonstrated splicing out of mRNA exon 4 [11]. Another study found a conserved mutation in the active site of ADA that ranges from codon 8 to 346 on Exon 9 with Arginine being replaced by Glutamine substitution causing changes to the hydropathy index has been hypothesized to change the deaminase activity of the enzyme [12]. 50 unrelated healthy patients also had their gene sequenced and none showed the specific conserved single nucleotide substitution found in carriers. Figure E (left) shows a confocal microscopic picture of immunostained ADA proteins (top-right) found in R1 and R2 subpopulations in Rabbit Gastric Mucosa ECL Cells.
The bottom-left image shows where A2BR protein and ADA merge on the cell surface of cells. Bottom-right shows colocalization of both proteins on the cell plasma membrane all studied by non-permeabilized cell preparations [14] |
Resources
[1] "Gene Ontology Database". Gene Ontology Consortium.
[2]Ontology Documentation http://geneontology.org/page/ontology-documentation
[3] Mosquera JL, Sánchez-Pla A "SerbGO: searching for the best GO tool". Nucleic Acids Research. 36 (Web Server issue): W368–71. doi:10.1093/nar/gkn256. PMC 2447766 . PMID 18480123. July 2008
[4] Holland M Steven, Rosenszweig D. Sergio “Immunodeficiencies” Infectious Diseases (Fourth Edition) 2017, Pages 705–722.e2 Volume 1
[5] ADA Gene https://ghr.nlm.nih.gov/gene/ADA
[6] Schrader WP, Stacy AR (Sep 1977). "Purification and subunit structure of adenosine deaminase from human kidney". The Journal of Biological Chemistry. 252 (18): 6409–6415. PMID 893413.
[7]”Summary: Adenosine/AMP deaminase” http://pfam.xfam.org/family/A_deaminase#Function
[8] “What is GO” https://www.ebi.ac.uk/training/online/course/goa-and-quickgo-quick-tour/what-go
[9] Cristalli G, Costanzi S, Lambertucci C, Lupidi G, Vittori S, Volpini R, Camaioni E (Mar 2001). "Adenosine deaminase: functional implications and different classes of inhibitors". Medicinal Research Reviews. 21 (2): 105–128. doi:10.1002/1098-1128(200103)21:2<105::AID-MED1002>3.0.CO;2-U. PMID 11223861.
[10] Wilson DK, Rudolph FB, Quiocho FA (May 1991). "Atomic structure of adenosine deaminase complexed with a transition-state analog: understanding catalysis and immunodeficiency mutations". Science. 252 (5010): 1278–1284. doi:10.1126/science.1925539. PMID 1925539.
[11] Akeson AL, Wiginton DA, States JC, Perme CM, Dusing MR, Hutton JJ.Mutations in the human adenosine deaminase gene that affect protein structure and RNA splicing.Proc Natl Acad Sci U S A. 1987 Aug;84(16):5947-51.
[12]Hellani Ali, Almasri, Nidal, Khaled K. Abu-Amero. A novel mutation in the ADA gene causing severe combined immunodeficiency in an Arab patient: A case report Journal of Medical Case Reports 3(1):6799 · April 2009
[13]