Defining The Similarities
Homology is a term coined from the greek word homologos, meaning: "relation".[1] It is used to describe shared ancestral traits between a pair of structures of one or more organism populations [1]. They can be a similar in structure and anatomical position but not necessarily in function between different organisms indicating a common ancestry or evolutionary origin. Homologies can range from similar skeletal limb structures among different species down to the same protein expression found in their genes. An example for instance is frogs, birds, rabbits and lizards all have different forelimbs, reflecting their different lifestyles. But those different forelimbs all share the same set of bones - the humerus, the radius, and the ulna. These are the same bones seen in fossils of the extinct transitional animal, Eusthenopteron, which demonstrates their common ancestry [5].
Homology is a term coined from the greek word homologos, meaning: "relation".[1] It is used to describe shared ancestral traits between a pair of structures of one or more organism populations [1]. They can be a similar in structure and anatomical position but not necessarily in function between different organisms indicating a common ancestry or evolutionary origin. Homologies can range from similar skeletal limb structures among different species down to the same protein expression found in their genes. An example for instance is frogs, birds, rabbits and lizards all have different forelimbs, reflecting their different lifestyles. But those different forelimbs all share the same set of bones - the humerus, the radius, and the ulna. These are the same bones seen in fossils of the extinct transitional animal, Eusthenopteron, which demonstrates their common ancestry [5].
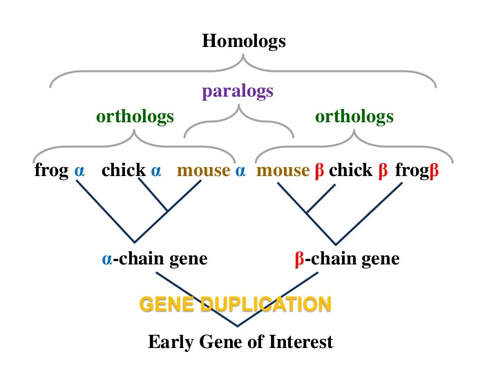
Orthologs, Paralogs, and Homologs
|
A key distinction in taxonomical organization and creation of phylogenetic linkages is identifying genes to be either homologs or paralogs of one another. Homology encumbers both orthologs and paralogs as laid out in figure on the left [8]. Orthologs are a result of speciation events as seen at the node of the α-chain gene. Paralogs arise from a duplication event as seen in the figure with the duplication of the gene interest into two different chain variations of the same gene, α-chain and β-chain [8].
|
ADA Gene Homologs in Common Organisms
|
White Cheeked Gibbon
(Nomascus leucogenys) ADA (Isoform X1) Accession ID: XP_003253658.1 Length: 363 aa 99% Identity |
|
Thale Cress (Arabidopsis thaliana)
Adenosine-deaminase like protein Accession ID: NP_192397.2 Length: 355 aa 29% Identity |
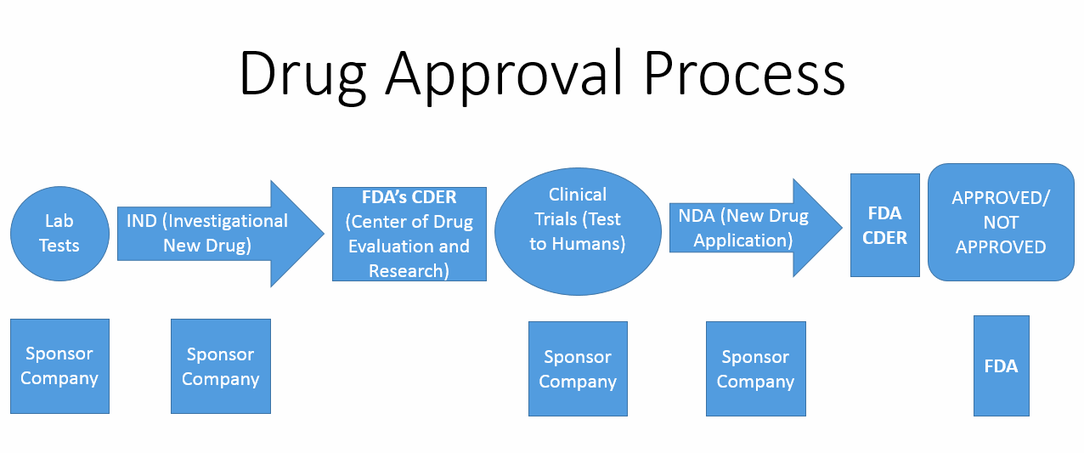
The drug approval flowchart above depicts the standardized process that scientist go through in the process of developing new drugs for diseases based on an extensive amount of previous studies & lab tests conducted on many non-human animals [6]
Applications/ Implementation
Evolutionary studies of homologs among different species allows us to better understand the role that genes and proteins of interest have on the cellular and organismal level. Conducting lab tests on taxonomically similar animals to humans with the same physiological, cellular, and biochemical responses to these genes as humans is crucial for therapy and drug development. Scientist are able to make accurate assessments of how to target these genes before proceeding to clinical trials.
Evolutionary studies of homologs among different species allows us to better understand the role that genes and proteins of interest have on the cellular and organismal level. Conducting lab tests on taxonomically similar animals to humans with the same physiological, cellular, and biochemical responses to these genes as humans is crucial for therapy and drug development. Scientist are able to make accurate assessments of how to target these genes before proceeding to clinical trials.
| protein_homologs.txt | |
| File Size: | 4 kb |
| File Type: | txt |
Resources
(1) Bower, Frederick Orpen (1906). "Plant Morphology". Congress of Arts and Science: Universal Exposition, St. Louis, 1904. Houghton, Mifflin. p. 64.
[2]Biology Online. "Homolgy" https://www.biology-online.org/dictionary/Homology
[3] BLAST ® » blastp suite » RID-96FCPX4F014
https://blast.ncbi.nlm.nih.gov/Blast.cgi#alnHdr_560960243
[4] Zimmer, Carl. Emlen, Douglas (2013) Evolution. 2nd Edition. New York, NY W. H. Freeman
[5] Lines of Evidence: The Science of Evolution. https://evolution.berkeley.edu/evolibrary/article/lines_04
[6] FDA's Drug Approval Process Flowchart. http://www.filipinoinvestor.com/2014/06/fdas-drug-approval-process-flow-chart.html 6/24/2016
[7] Jensen A. Roy "Orthologs and Paralos- we need to get it right"https://www.ncbi.nlm.nih.gov/pmc/articles/PMC138949/
[8] https://biology.stackexchange.com/questions/4962/what-is-the-difference-between-orthologs-paralogs-and-homologs
(1) Bower, Frederick Orpen (1906). "Plant Morphology". Congress of Arts and Science: Universal Exposition, St. Louis, 1904. Houghton, Mifflin. p. 64.
[2]Biology Online. "Homolgy" https://www.biology-online.org/dictionary/Homology
[3] BLAST ® » blastp suite » RID-96FCPX4F014
https://blast.ncbi.nlm.nih.gov/Blast.cgi#alnHdr_560960243
[4] Zimmer, Carl. Emlen, Douglas (2013) Evolution. 2nd Edition. New York, NY W. H. Freeman
[5] Lines of Evidence: The Science of Evolution. https://evolution.berkeley.edu/evolibrary/article/lines_04
[6] FDA's Drug Approval Process Flowchart. http://www.filipinoinvestor.com/2014/06/fdas-drug-approval-process-flow-chart.html 6/24/2016
[7] Jensen A. Roy "Orthologs and Paralos- we need to get it right"https://www.ncbi.nlm.nih.gov/pmc/articles/PMC138949/
[8] https://biology.stackexchange.com/questions/4962/what-is-the-difference-between-orthologs-paralogs-and-homologs