Post-Translational Modifications
|
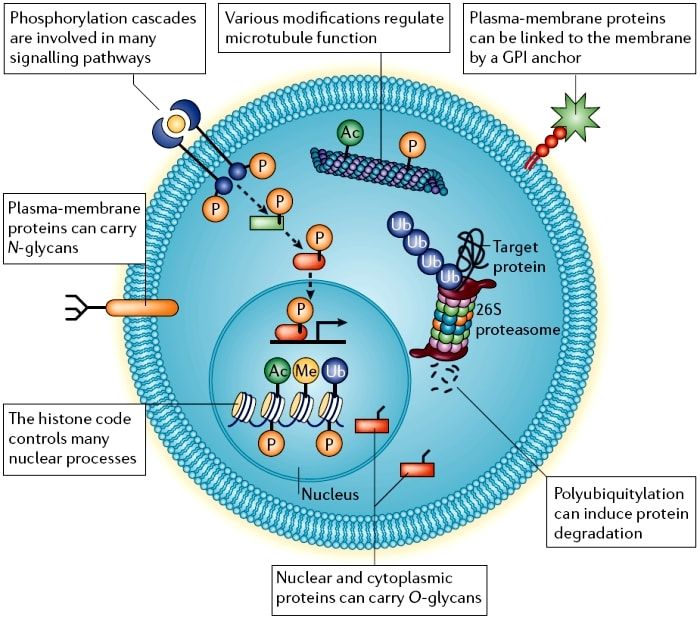
Diagram A shows cellular post translational modifications with inclusion of various cellular components involved in different protein modifying pathways[3].
|
The proteome is the entire set of proteins that are encoded and translated by the genome. The trajectory to the protein’s determinant function can be chemically altered through various mechanisms called post translational modifications (PTM). These protein modifications occur through multiple possible reactions like phosphorylation, methylation, acetylation, glycosylation, nitrosylation, ubiquitination, lipidation, and proteolysis. These modifications play a key role in protein function as they regulate protein activity, localization, and interaction with other like other proteins, nucleic acids, lipids and cofactors [1]. PTM’s occurs at amino acid side chains or peptide linkages which are mediated by enzyme activity. Proteomics, the study of the proteome, allows for studying detailed information about the molecular biology of the living cell and the organism during disease development[4]. |
|
Mechanism of Study
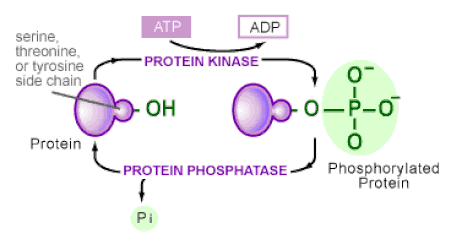
For my specific disease I will focus on phosphorylation sites found in ADA. This mechanism is known to add a hydrophilic group to an amino acid chain that alters the interactions with nearby amino acids. These modified interactions can have a pivotal role how well ADA performs to signal cell growth and replication. Learning more on the differences in phosphorylation of ADA in healthy individuals vs ADA deficient patients will help identify possible previously unknown changes to the protein conformation and how to prevent them all together. |
Diagram B shows the mechanism of phosphorylation and the attachment of a phosphate group to replace the hydroxyl group found on the amino acid side chain.
|
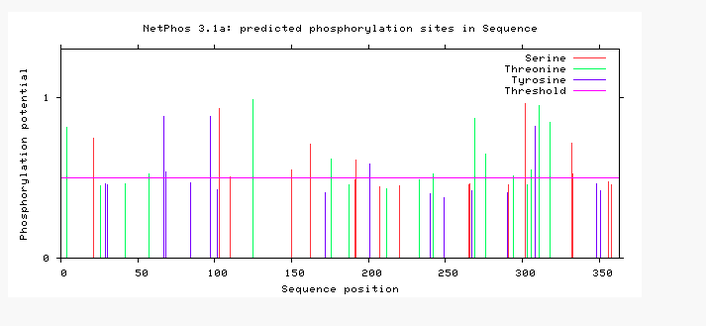
Phosphorylation predictions in ADA1
With the use of the NetPhos program, predictions were made on the human isoform of ADA1 identifying possible amino acid sites susceptible to phosphorylation. The prediction algorithm predicted a total of 46 amino acid positions where phosphorylation was likely to occur.
Results
These potential phosphorylation sites identified through NetPhos show positive signs possible post-translational modifications occurring to Adenosine Deaminase. To confirm just how significant the PTM changes are among ADA deficient mice, I will transition to conducting a stable isotope amino acid labeling experiment followed up by an HLC-Mass spectometry analysis. This will allow better understanding of just how much phosphorylation plays a role in the metabolic cascade pathway that affects pulmonary function.
These potential phosphorylation sites identified through NetPhos show positive signs possible post-translational modifications occurring to Adenosine Deaminase. To confirm just how significant the PTM changes are among ADA deficient mice, I will transition to conducting a stable isotope amino acid labeling experiment followed up by an HLC-Mass spectometry analysis. This will allow better understanding of just how much phosphorylation plays a role in the metabolic cascade pathway that affects pulmonary function.
Resources
1) Overview of Post-Translational Modifications (PTMs) https://www.thermofisher.com/us/en/home/life-science/protein-biology/protein-biology-learning-center/protein-biology-resource-library/pierce-protein-methods/overview-post-translational-modification.html
2) Posttranslational Modification A. Bürkle, in Encyclopedia of Genetics, 2001
3) The Proteome: Discovering the Structure and Function of ProteinsBy: Jill Adams, Ph.D. (Freelance science writer in Albany, NY) © 2008 Nature Education Citation: Adams, J. (2008) The Proteome: Discovering the Structure and Function of Proteins. Nature Education 1(3):6
4) What is proteome analysis? http://www.sdu.dk/Nat/CPA/proteomics.html
1) Overview of Post-Translational Modifications (PTMs) https://www.thermofisher.com/us/en/home/life-science/protein-biology/protein-biology-learning-center/protein-biology-resource-library/pierce-protein-methods/overview-post-translational-modification.html
2) Posttranslational Modification A. Bürkle, in Encyclopedia of Genetics, 2001
3) The Proteome: Discovering the Structure and Function of ProteinsBy: Jill Adams, Ph.D. (Freelance science writer in Albany, NY) © 2008 Nature Education Citation: Adams, J. (2008) The Proteome: Discovering the Structure and Function of Proteins. Nature Education 1(3):6
4) What is proteome analysis? http://www.sdu.dk/Nat/CPA/proteomics.html