What are protein networks?
Protein networks display the interactions among various proteins that are involved in the same pathway or cellular function. Protein-Protein interactions (PPI’s) are mathematical representations of the of the physical contact between proteins. An all encompassing graphical representations of this network can be constructed in to an interactome. The interactome of an organism is computed through prediction algorithms extracted from molecular databases. Gaining information about the molecular processes can be done through 2 methods: computational or experimental. Regardless of the approach taken there is no perfect method. Experimental methods are typically reinforced through computational algorithms to create a more comprehensive global view of the protein network. Understanding protein-protein interactions is crucial for understanding the physiological characteristics of the cell in both diseased and normal states [1].
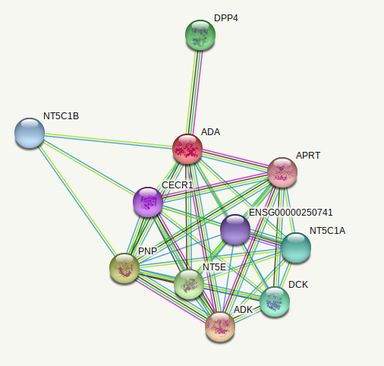
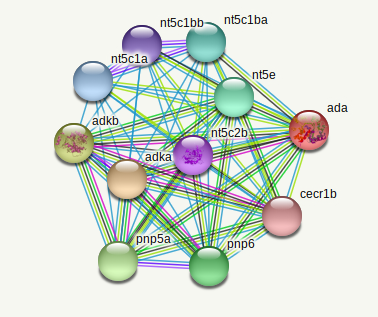
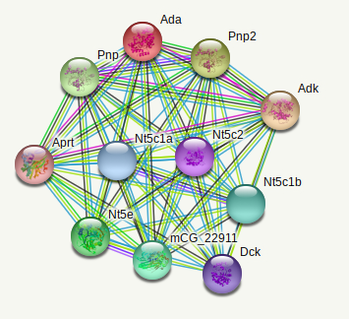
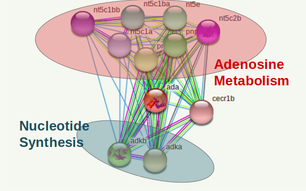
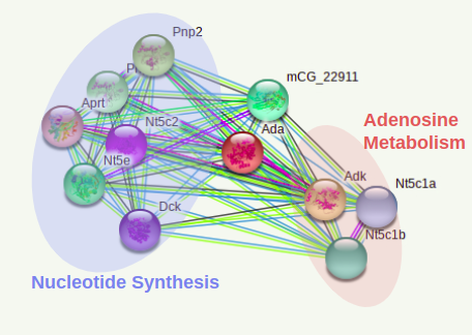
The 2 ADA interactomes below were done with the PPI database of the online tool STRING [2].
Protein networks display the interactions among various proteins that are involved in the same pathway or cellular function. Protein-Protein interactions (PPI’s) are mathematical representations of the of the physical contact between proteins. An all encompassing graphical representations of this network can be constructed in to an interactome. The interactome of an organism is computed through prediction algorithms extracted from molecular databases. Gaining information about the molecular processes can be done through 2 methods: computational or experimental. Regardless of the approach taken there is no perfect method. Experimental methods are typically reinforced through computational algorithms to create a more comprehensive global view of the protein network. Understanding protein-protein interactions is crucial for understanding the physiological characteristics of the cell in both diseased and normal states [1].
The 2 ADA interactomes below were done with the PPI database of the online tool STRING [2].
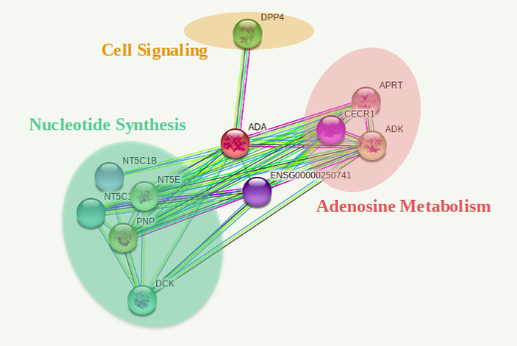
Cellular Process Network Comparisons
The interactome of both species appear highly conserved among cellular processes as well as specific proteins involved in each pathway. It is definitive to conclude the gene function is uniform among both homologs.
RESULTS
The protein interactions found by STRING appear consistent with previous studies on the metabolic processes. An interesting find that could lead to more specific protein studies is CECR1 found in the human and zebrafish isoform involved in adenosine degradation. This gene is encoded in the cat eye syndrome chromosome location of the genome involved in the pathway responsible for other physiological defects possibly even pulmonary defects.
The protein interactions found by STRING appear consistent with previous studies on the metabolic processes. An interesting find that could lead to more specific protein studies is CECR1 found in the human and zebrafish isoform involved in adenosine degradation. This gene is encoded in the cat eye syndrome chromosome location of the genome involved in the pathway responsible for other physiological defects possibly even pulmonary defects.
Resources:
1) Protein- Protein Interaction Networks https://www.ebi.ac.uk/training/online/course/network-analysis-protein-interaction-data-introduction/protein-protein-interaction-networks
2) STRING https://string-db.org/
1) Protein- Protein Interaction Networks https://www.ebi.ac.uk/training/online/course/network-analysis-protein-interaction-data-introduction/protein-protein-interaction-networks
2) STRING https://string-db.org/